Electron Cryomicroscopy
Lead Researcher: Marcus A. Brubaker
The structure of biological molecules like proteins and viruses play a critical role in understanding life at a cellular level and are important to the discovery of new medical treatments. However determining these structures is a difficult task due to their small size and delicate biological nature. Electron Cryomicroscopy (cryo-EM for short) is an emerging new technique for structure determination which uses large numbers of low-contrast, high-noise 2D images to attempt to estimate 3D structure.
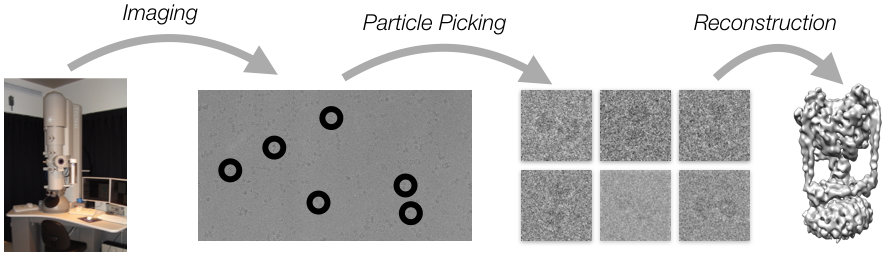
This project aims to explore the computational aspects of this reconstruction and estimation problem. This includes a range of problems such as particle picking [Zhao et al. 2013], 3D reconstruction [Punjani et al. 2017; Punjani et al. 2016], image clustering, structural clustering, deconvolution [Zhao et al. 2015], motion estimation and correction [Rubinstein and Brubaker 2015] and others.

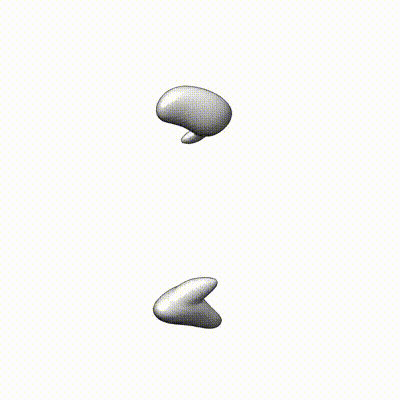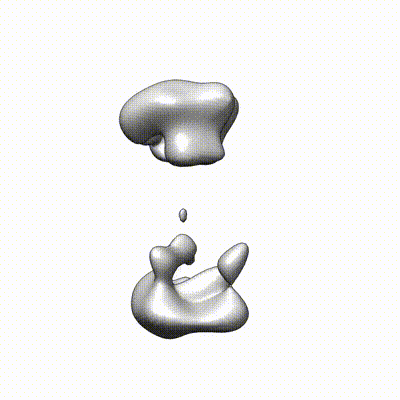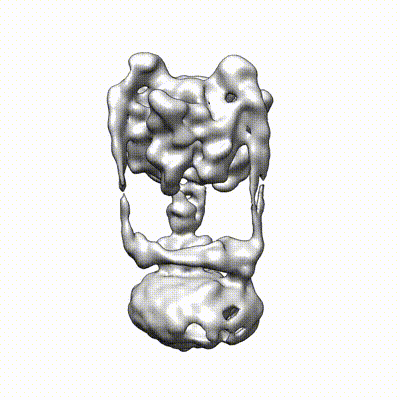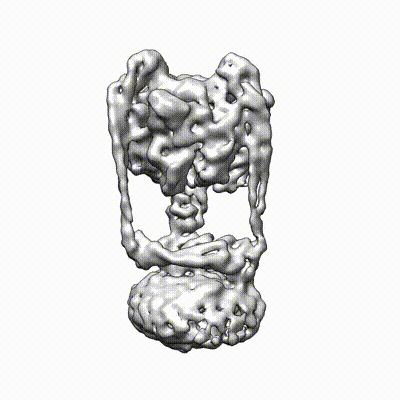
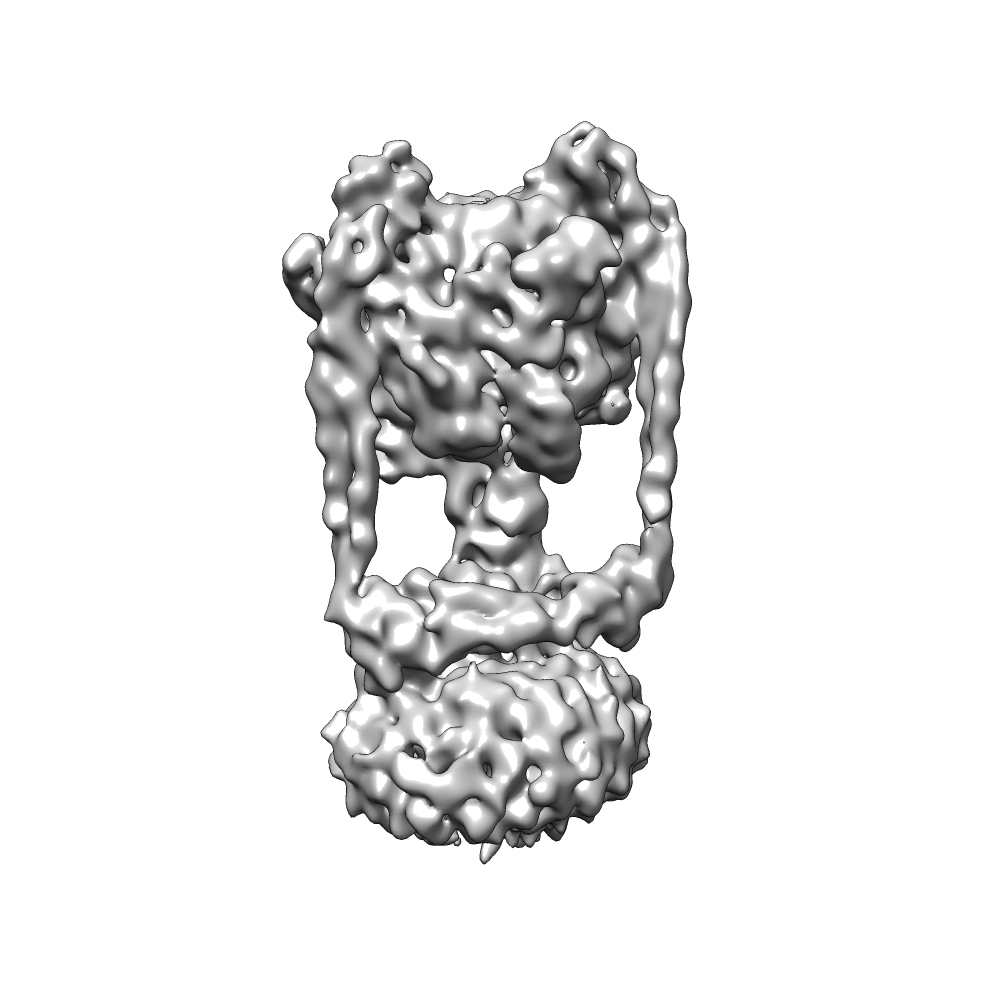
The following is a selected list of publications related to this project:
- Zhao, J., Brubaker, M.A., and Rubinstein, J.L. 2013. TMaCS: A hybrid template matching and classification system for partially-automated particle selection. Journal of Structural Biology 181, 3, 234–242.
- Punjani, A., Rubinstein, J.L., Fleet, D.J., and Brubaker, M.A. 2017. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nature Methods 14, 3, 290–296.
- Punjani, A., Brubaker, M.A., and Fleet, D.J. 2016. Building Proteins in a Day: Efficient 3D Molecular Structure Estimation with Electron Cryomicroscopy. IEEE Transactions on Pattern Analysis and Machine Intelligence.
- Zhao, J., Brubaker, M.A., Benlekbir, S., and Rubinstein, J.L. 2015. Description and comparison of algorithms for correcting anisotropic magnification in cryo-EM images. Journal of Structural Biology 192, 2, 209–215.
- Rubinstein, J.L. and Brubaker, M.A. 2015. Alignment of cryo-EM movies of individual particles by optimization of image translations. Journal of Structural Biology 192, 2, 188–195.